Tryptophan synthase
Tryptophan synthase
Jump to navigation
Jump to search
| Tryptophan Synthase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
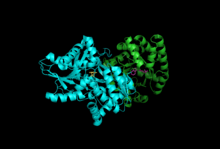 Subunits: Beta Subunit, Alpha Subunit with PLP, IGP | |||||||||
| Identifiers | |||||||||
| EC number | 4.2.1.20 | ||||||||
| CAS number | 9014-52-2 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
Tryptophan synthase or tryptophan synthetase is an enzyme that catalyzes the final two steps in the biosynthesis of tryptophan.[1] It is commonly found in Eubacteria,[2]Archaebacteria,[3]Protista,[4]Fungi,[5] and Plantae.[6] However, it is absent from Animalia.[7] It is typically found as an α2β2 tetramer. The α subunits catalyze the reversible formation of indole and glyceraldehyde-3-phosphate (G3P) from indole-3-glycerol phosphate (IGP). The β subunits catalyze the irreversible condensation of indole and serine to form tryptophan in a pyridoxal phosphate (PLP) dependent reaction. Each α active site is connected to a β active site by a 25 angstrom long hydrophobic channel contained within the enzyme. This facilitates the diffusion of indole formed at α active sites directly to β active sites in a process known as substrate channeling.[8] The active sites of tryptophan synthase are allosterically coupled.[9]
Contents
1 Enzyme structure
2 Enzyme mechanism
3 Biological function
4 Substrate scope
5 Disease relevance
6 Evolution
7 Historical significance
8 See also
9 References
Enzyme structure[edit]
Subunits: Tryptophan synthase typically exists as an α-ββ-α complex. The α and β subunits have molecular masses of 27 and 43 kDa respectively. The α subunit has a TIM barrel conformation. The β subunit has a fold type II conformation and a binding site adjacent to the active site for monovalent cations.[10] Their assembly into a complex leads to structural changes in both subunits resulting in reciprocal activation. There are two main mechanisms for intersubunit communication. First, the COMM domain of the β-subunit and the α-loop2 of the α-subunit interact. Additionally, there are interactions between the αGly181 and βSer178 residues.[11] The active sites are regulated allosterically and undergo transitions between open, inactive, and closed, active, states.[9]
Indole-3-glycerol binding site: See image 1.
Indole and serine binding site: See image 1.
Hydrophobic channel: The α and β active sites are separated by a 25 angstrom long hydrophobic channel contained within the enzyme allowing for the diffusion of indole. If the channel did not exist, the indole formed at an α active site would quickly diffuse away and be lost to the cell as it is hydrophobic and can easily cross membranes. As such, the channel is essential for enzyme complex function.[12]

Image 2: Proposed mechanism of tryptophan synthase
Enzyme mechanism[edit]
α subunit reaction: The α subunit catalyzes the formation of indole and G3P from a retro-aldol cleavage of IGP. The αGlu49 and αAsp60 are thought to be directly involved in the catalysis as shown.[8] The rate limiting step is the isomerization of IGP.[13] See image 2.
β subunit reaction: The β subunit catalyzes the β-replacement reaction in which indole and serine condense to form tryptophan in a PLP dependent reaction. The βLys87, βGlu109, and βSer377 are thought to be directly involved in the catalysis as shown.[8] Again, the exact mechanism has not been conclusively determined. See image 2.
Net reaction: See image 3.

Image 3: Reaction catalyzed by tryptophan synthase

Image 1: Active sites for α and β subunits showing hypothesized catalytic residues
Biological function[edit]
Tryptophan synthase is commonly found in Eubacteria, Archaebacteria, Protista, Fungi, and Plantae. It is absent from animals such as humans. Tryptophan is one of the twenty standard amino acids and one of nine essential amino acids for humans. As such, tryptophan is a necessary component of the human diet.
Substrate scope[edit]
Tryptophan synthetase is also known to accept indole analogues, e.g., fluorinated or methylated indoles, as substrates, generating the corresponding tryptophan analogues.[14]
Disease relevance[edit]
As humans do not have tryptophan synthase, this enzyme has been explored as a potential drug target.[15] However, it is thought that bacteria have alternate mechanisms to produce amino acids which might make this approach less effective. In either case, even if the drug only weakens bacteria, it might still be useful as the bacteria are already vulnerable in the hostile host environment. As such, the inhibition of tryptophan synthase along with other PLP-enzymes in amino acid metabolism has the potential to help solve medical problems.[16]
Inhibition of tryptophan synthase and other PLP-enzymes in amino acid metabolism has been suggested for:
- Treatment of tuberculosis[15]
- Treatment of ocular and genital infections[17]
- Treatment of cryptosporidiosis[15]
Herbicide use[18]
Evolution[edit]
It is thought that early in evolution the trpB2 gene was duplicated. One copy entered the trp operon as trpB2i allowing for its expression with trpA. TrpB2i formed transient complexes with TrpA and in the process activated TrpA unidirectionally. The other copy remained outside as trpB2o, and fulfilled an existing role or played a new one such as acting as a salvage protein for indole. TrpB2i evolved into TrpB1, which formed permanent complexes with trpA resulting in bidirectional activation. The advantage of the indole salvage protein declined and the TrpB gene was lost. Finally, the TrpB1 and TrpA genes were fused resulting in the formation the bifunctional enzyme.[19]
Historical significance[edit]
Tryptophan synthase was the first enzyme identified that had two catalytic capabilities that were extensively studied. It was also the first identified to utilize substrate channeling. As such, this enzyme has been studied extensively and is the subject of great interest.[8]
See also[edit]
- Lyase
- Synthase
- Tryptophan synthase (indole-salvaging)
References[edit]
^ Dunn MF, Niks D, Ngo H, Barends TR, Schlichting I (June 2008). "Tryptophan synthase: the workings of a channeling nanomachine". Trends in Biochemical Sciences. 33 (6): 254–64. doi:10.1016/j.tibs.2008.04.008. PMID 18486479..mw-parser-output cite.citation{font-style:inherit}.mw-parser-output q{quotes:"""""""'""'"}.mw-parser-output code.cs1-code{color:inherit;background:inherit;border:inherit;padding:inherit}.mw-parser-output .cs1-lock-free a{background:url("//upload.wikimedia.org/wikipedia/commons/thumb/6/65/Lock-green.svg/9px-Lock-green.svg.png")no-repeat;background-position:right .1em center}.mw-parser-output .cs1-lock-limited a,.mw-parser-output .cs1-lock-registration a{background:url("//upload.wikimedia.org/wikipedia/commons/thumb/d/d6/Lock-gray-alt-2.svg/9px-Lock-gray-alt-2.svg.png")no-repeat;background-position:right .1em center}.mw-parser-output .cs1-lock-subscription a{background:url("//upload.wikimedia.org/wikipedia/commons/thumb/a/aa/Lock-red-alt-2.svg/9px-Lock-red-alt-2.svg.png")no-repeat;background-position:right .1em center}.mw-parser-output .cs1-subscription,.mw-parser-output .cs1-registration{color:#555}.mw-parser-output .cs1-subscription span,.mw-parser-output .cs1-registration span{border-bottom:1px dotted;cursor:help}.mw-parser-output .cs1-hidden-error{display:none;font-size:100%}.mw-parser-output .cs1-visible-error{font-size:100%}.mw-parser-output .cs1-subscription,.mw-parser-output .cs1-registration,.mw-parser-output .cs1-format{font-size:95%}.mw-parser-output .cs1-kern-left,.mw-parser-output .cs1-kern-wl-left{padding-left:0.2em}.mw-parser-output .cs1-kern-right,.mw-parser-output .cs1-kern-wl-right{padding-right:0.2em}
^ Jablonski P, Jablonski L, Pintado O, Sriranganathan N, Howde C (September 1996). "Tryptophan synthase: Identification of Pasteurella multocida tryptophan synthase B-subunit by antisera against strain PI059". Microbiology. 142: 115–21. doi:10.1099/13500872-142-1-115. PMID 8581158.
^ Lazcano A, Diaz-Villgomez E, Mills T, Oro J (March 1995). "On the levels of enzymatic substrate specificity: Implications for the early evolution of metabolic pathways". Advances in Space Research. 15 (3): 345–56. doi:10.1016/S0273-1177(99)80106-9. PMID 11539248.
^ Anderson I, Watkins R, Samuelson J, Spencer D, Majoros W, Grey M, Loftus B (August 2005). "Gene Discovery in the Acanthamoeba castellanii Genome". Protist. 156 (2): 203–14. doi:10.1016/j.protis.2005.04.001. PMID 16171187.
^ ,Ireland C, Peekhaus N, Lu P, Sangari R, Zhang A, Masurekar P, An Z (April 2008). "The tryptophan synthetase gene TRP1 of Nodulisporium sp.: molecular characterization and its relation to nodulisporic acid A production". Appl Microbiol Biotechnol. 79 (3): 451–9. doi:10.1007/s00253-008-1440-3. PMID 18389234.
^ Sanjaya, Hsiao PY, Su RC, Ko SS, Tong CG, Yang RY, Chan MT (April 2008). "Overexpression of Arabidopsis thaliana tryptophan synthase beta 1 (AtTSB1) in Arabidopsis and tomato confers tolerance to cadmium stress". Plant Cell Environ. 31 (8): 1074–85. doi:10.1111/j.1365-3040.2008.01819.x. PMID 18419734.CS1 maint: Multiple names: authors list (link)
^ Eckert SC, Kubler E, Hoffmann B, Braus GH (June 2000). "The tryptophan synthase-encoding trpB gene of Aspergillus nidulans is regulated by the cross-pathway control system". Mol Gen Genet. 263 (5): 867–76. doi:10.1007/s004380000250. PMID 10905354.
^ abcd Raboni S, Bettati S, Mozzarelli A (April 2009). "Tryptophan synthase: a mine for enzymologists". Cell Mol Life Sci. 66 (14): 2391–403. doi:10.1007/s00018-009-0028-0. PMID 19387555.
^ ab Fatmi MQ, Ai R, Chang CA (September 2009). "Synergistic regulation and ligand-induced conformational changes of tryptophan synthase". Biochemistry. 48 (41): 9921–31. doi:10.1021/bi901358j. PMID 19764814.
^ Grishin NV, Phillips MA, Goldsmith EJ (July 1995). "Modeling of the spatial structure of ornithine decarboxylases". Protein Sci. 4 (7): 1291–304. doi:10.1002/pro.5560040705. PMC 2143167. PMID 7670372.
^ Schneider TR, Gerhardt E, Lee M, Liang PH, Anderson KS, Schlichting I (April 1998). "Loop closure and intersubunit communication in tryptophan synthase". Biochemistry. 37 (16): 5394–406. doi:10.1021/bi9728957. PMID 9548921.
^ Huang X, Holden HM, Raushel FM (2001). "Channeling of Substrates and Intermediates in Enzyme-Catalyzes Reactions". Annu Rev Biochem. 70: 149–80. doi:10.1146/annurev.biochem.70.1.149. PMID 11395405.
^ Anderson KS, Miles EW, Johnson KA (May 1991). "Serine modulates substrate channeling in tryptophan synthase. A novel intersubunit triggering mechanism". J Biol Chem. 266 (13): 8020–33. PMID 1902468.
^ Wilcox, Michael (1974-06-01). "The enzymatic synthesis of l-tryptophan analogues". Analytical Biochemistry. 59 (2): 436–440. doi:10.1016/0003-2697(74)90296-6.
^ abc Chaudhary K, Roos DS (September 2005). "Protozoan genomics for drug discovery". Nat Biotechnol. 23 (9): 1089–91. doi:10.1038/nbt0905-1089. PMID 16151400.
^ Becker D, Selbach M, Rollenhagen C, Ballmaier M, Meyer TF, Mann M, Bumann D (March 2006). "Robust Salmonella metabolism limits possibilities for new antimicrobials". Nature. 440 (7082): 303–7. doi:10.1038/nature04616. PMID 16541065.
^ Caldwell HD, Wood H, Crane D, Baily R (June 2003). "Polymorphisms in Chlamydia trachomatis tryptophan synthase genes differentiate between genital an ocular isolates". J Clin Invest. 111 (11): 1757–69. doi:10.1172/JCI17993. PMC 156111. PMID 12782678.
^ Kulik V, Hartmann E, Weyand M, Frey M, Gierl A, Niks D, Dunn MF, Schlichting I (September 2005). "On the structural basis of the catalytic mechanism and the regulation of the α-subunit of tryptophan synthase from Salmonella typhimurium and BXI from maize, two evolutionarily related enzymes". J Mol Biol. 352 (3): 608–20. doi:10.1016/j.jmb.2005.07.014. PMID 16120446.
^ Leopoldseder S, Hettwer S, Sterner R (November 2006). "Evolution of Multi-Enzyme Complexes: The Case of Tryptophan Synthase". Biochemistry. 45 (47): 14111–9. doi:10.1021/bi061684b. PMID 17115706.
Categories:
- EC 4.2.1
(window.RLQ=window.RLQ||).push(function(){mw.config.set({"wgPageParseReport":{"limitreport":{"cputime":"0.472","walltime":"0.555","ppvisitednodes":{"value":1473,"limit":1000000},"ppgeneratednodes":{"value":0,"limit":1500000},"postexpandincludesize":{"value":89831,"limit":2097152},"templateargumentsize":{"value":1509,"limit":2097152},"expansiondepth":{"value":10,"limit":40},"expensivefunctioncount":{"value":2,"limit":500},"unstrip-depth":{"value":1,"limit":20},"unstrip-size":{"value":58476,"limit":5000000},"entityaccesscount":{"value":2,"limit":400},"timingprofile":["100.00% 462.418 1 -total"," 68.75% 317.926 1 Template:Reflist"," 60.93% 281.730 19 Template:Cite_journal"," 16.66% 77.028 2 Template:Infobox"," 15.85% 73.301 1 Template:Enzyme"," 5.14% 23.763 3 Template:Navbox"," 5.11% 23.629 1 Template:Portal_bar"," 3.56% 16.442 1 Template:Multienzyme_complexes"," 1.87% 8.637 1 Template:Enzymes"," 1.68% 7.787 1 Template:Carbon-oxygen_lyases"]},"scribunto":{"limitreport-timeusage":{"value":"0.291","limit":"10.000"},"limitreport-memusage":{"value":4234998,"limit":52428800}},"cachereport":{"origin":"mw1320","timestamp":"20181118062714","ttl":1900800,"transientcontent":false}}});mw.config.set({"wgBackendResponseTime":114,"wgHostname":"mw1323"});});